 Producenci
Producenci

Structural Multiplex cfDNA Reference Standard
 Opis
Opis
Format: cfDNA
The Structural Multiplex cfDNA Reference Standard provides biologically relevant quality control material, which can be used to assess the performance of NGS ctDNA assays that detect complex structural variants.
With this product you can:
• Evaluate the effect of genomic context on variant detection in ctDNA samples
• Analyze the robustness of your bioinformatics pipeline
• Optimize and validate new ctDNA workflows for structural variant detection
• Compare performance of fragmented to high molecular weight reference standards
Structural variants including gene translocations, fusions, copy number variants (CNV) and large INDELs are commonly found in tumours, and can serve as clinical biomarkers to identify cancer, to monitor cancer treatment effectiveness and disease progression. CNVs such as amplification of MET in combination with mutations in EGFR (T790M) can lead to Gefitinib resistance in lung cancer 1, 2.
PCR-based assays3, 4 and NGS4 that better detect structural variants are constantly emerging. However, the detection of structural variants presents several challenges and bioinformatics pipelines are continuously evolving to improve confidence in structural variant detection.
The Structural Multiplex cfDNA Reference Standard covers a wide range of mutations in a defined genomic context. This standard consists of DNA derived from human cell lines that is fragmented to an average size of 160 bp to closely resemble cfDNA extracted from human plasma.
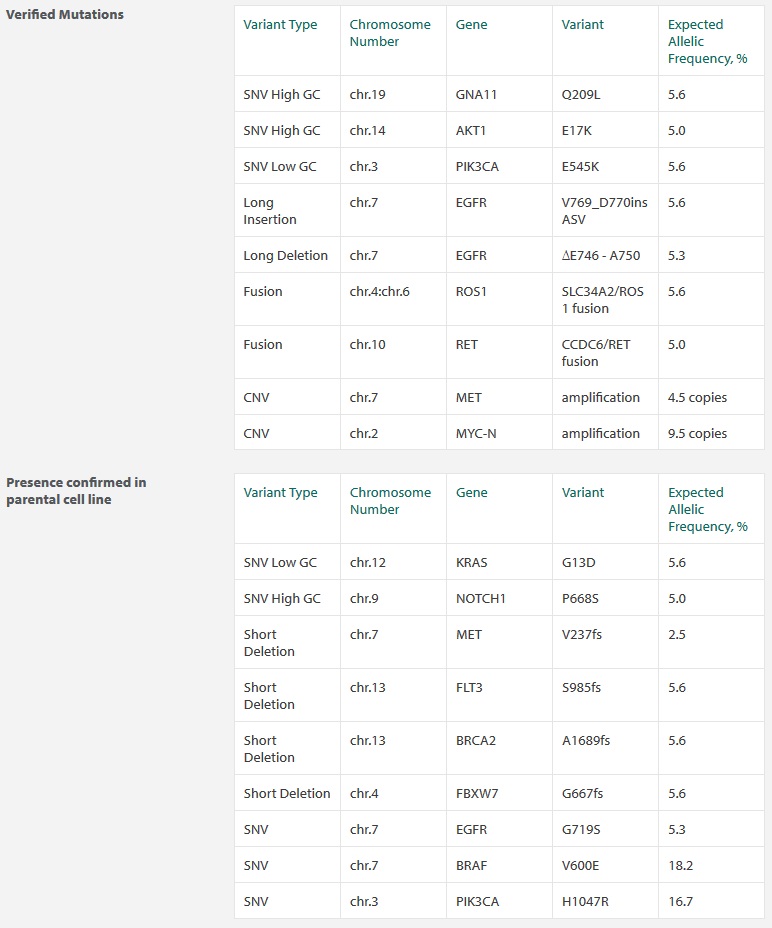
This product is designed to challenge your molecular and bioinformatic work flow by providing validated copy number variants/amplifications, translocations, and large insertions/deletions. Additionally, you may examine the genomic context of variants within regions of specific GC-content (high vs. low). The Structural Multiplex cfDNA Reference Standard includes 9 variants validated by ddPCR, with most of them centred at 5% allelic frequency.
Highlight features of the Structural Multiplex include RET and ROS1 fusion variants, MYC-N and MET focal amplifications and a BRCA2 variant.
The Structural Multiplex is also available in high molecular weight gDNA (HD753) and FFPE (HD789) format.
References:
- Engelman JA, Zejnullahu K, Mitsudomi T, Song Y, Hyland C, Park JO, et al. MET amplification leads to gefitinib resistance in lung cancer by activating ERBB3 signaling. Science 2007,316:1039–43
- Bean J, Brennan C, Shih JY, Riely G, Viale A, Wang L, et al. MET amplification occurs with or without T790M mutations in EGFR mutant lung tumors with acquired resistance o gefitinib or erlotinib. Proc Natl Acad Sci.2007,104(52):20932–7
- Olsson, E.; Winter, C. et al. Serial monitoring of circulating tumor DNA in patients with primary breast cancer for detection of occult metastatic disease. EMBO Molec. Med. 2015, 7(12), 1034–47.
- Reinert, T.; Schøler, L.V. et al. Analysis of circulating tumour DNA to monitor disease burden following colorectal cancer surgery. Gut 2015, 0:1–10.
Technical Data
Genes Covered: AKT1, BRCA2, EGFR, FBXW7, FLT3, GNA11, KRAS, MET, MYC, NOTCH1, PIK3CA, RET, ROS1
Cosmics: 1,500+
Allelic Frequencies: 4.8-5.6% and 4.5 & 9.5 x amplification for the CNV
Buffer: Tris-EDTA (10mM Tris-HCl, 1mM EDTA), pH 8.0
Product Information
Fragment Size: 160 bp
Unit Size: 350 ng per vial
Concentration: 20 ng/µl
General Information
Storage: 4°C
Expiry: 12 months from the date of manufacture
Cell Line Background: Please contact for more information
Quality Control
Fragmentation Size: D1000 DNA ScreenTape assay
Allelic Frequency: Droplet Digital™ PCR
Quantification: Qubit dsDNA BR Assay (post-fragmentation)

